URL: https://www.desy.de/news/news_search/index_eng.html
Breadcrumb Navigation
DESY News: Unravelling the secrets of the malaria parasite
News
News from the DESY research centre
Unravelling the secrets of the malaria parasite
For the first time, scientists have identified a lipocalin protein in the malaria parasite Plasmodium falciparum. The discovery helps to better understand the life cycle of the parasite that is a major health burden in large parts of the world. The cooperation between the groups of Tim Gilberger from the Centre for Structural Systems Biology CSSB (Cellular Parasitology Department at Bernhard Nocht Institute for Tropical Medicine/ Universität Hamburg) at DESY and Matthias Wilmanns from the Hamburg branch of the European Molecular Biology Laboratory EMBL describes the discovery in the journal Cell Reports. CSSB is a cooperation of nine institutions, including DESY, that have deputed scientists to the centre.

Lipocalins are a family of funnel-like proteins that channel insoluble molecules such as lipids and steroids into cells. They are present in almost every organism, associated with numerous functions and show an extreme diversity that makes them hard to identify. There were no lipocalins identified in P. falciparum. Gilberger's group focused on a single gene in the malaria parasite that was predicted to encode a protein with a lipocalin superfamily signature.
To validate the identified protein the group teamed up with Willmann’s group at EMBL Hamburg. “Lipocalins don’t have a conserved primary structure but all members of the family have a similar crystal structure,” explains EMBL's Thomas Crosskey, a co-author of the paper. “Our analysis of the protein using X-ray crystallography at the PETRA III synchrotron’s P13 beamline revealed a tetrameric structure of monomers with a classic lipocalin fold thus confirming the protein as a lipocalin.” It was named P. falciparum lipocalin (PfLCN).
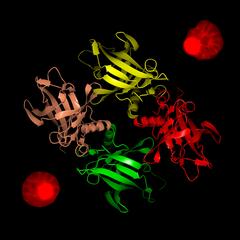
After identifying, confirming the structure and locating PfLCN, the scientists moved on to investigate the protein’s function within the malaria parasite. “First we needed to find out whether PfLCN plays an important role in blood stage proliferation of the parasite,” noted Burda. “To accomplish this we removed PfLCN from the parasite and observed that without the lipocalin the parasite could not replicate in its host cell anymore.”
Intrigued by its localization to the vacuolar compartments that represent intracellular interfaces vulnerable to oxidative damage, the scientists then exposed the modified parasite to a radical scavenging molecule. “We were surprised to discover that this resulted in a partial rescue,” explains Gilberger “this suggest that one important function of PfLCN might be to play a role in reducing oxidative cell damage in the malaria parasite.” Next, Gilberger's group plans to investigate what PfLCN binds to and how it interacts with other molecules such as lipids or other hydrophobic compounds.
This collaboration not only solved a small piece of the mystery surrounding how P. falciparum ensures its survival in the host’s red blood cells but also demonstrated the success of using a combination of complementary methods to generate a fuller understanding of a protein’s structure and function. “CSSB was established to allow scientist to combine the powers of X-ray crystallography at PETRA III with complimentary microscopic techniques to answer important biological questions surrounding infection,” explains Wilmanns who was CSSB’s founding director.
Reference:
Structure-Based Identification and Functional Characterization of a Lipocalin in the Malaria Parasite Plasmodium falciparum; Paul-Christian Burda, Thomas Crosskey, Katharina Lauk, Aimo Zurborg, Christoph Söhnchen, Benjamin Liffner, Louisa Wilcke, Emma Pietsch, Jan Strauss, Cy M. Jeffries, Dmitri I. Svergun, Danny W. Wilson, Matthias Wilmanns, Tim-Wolf Gilberger; Cell Reports, 2020; DOI: 10.1016/j.celrep.2020.107817



